Artificial intelligence agents are emerging as powerful applications of large language models (LLMs), automating complex tasks and enabling scientific data exploration. However, their use in biomedical data analysis remains limited by the difficulty of handling specialized tools and multistep reasoning. Here we introduce BioMedAgent, a self-evolving LLM multi-agent framework, which learns to use diverse bioinformatics tools and chain them into executable workflows through interactive exploration and memory retrieval algorithms. It allows biomedical users to initiate tasks using natural language, without requiring computational expertise. Evaluated on our newly released BioMed-AQA benchmark comprising 327 biomedical data tasks, BioMedAgent achieved a 77% success rate, surpassing other LLM agents, and generalized robustly to the external BixBench dataset. Beyond benchmarks, it autonomously performs cross-omics analysis, machine-learning modelling and pathology image segmentation, highlighting its potential to advance biomedical research and extend to other scientific domains requiring complex tool integration and multistep reasoning.
The biomedical field is experiencing rapid data expansion, driven by emerging technologies that generate vast amounts of medical texts, images and omics data1,2. The widespread adoption of electronic health records globally has greatly facilitated the collection of clini-cal data on diagnostics, treatments and follow-ups3,4. Advancements in digital pathology and high-resolution imaging technologies, such as whole-slide imaging, magnetic resonance imaging and computed tomography, have revolutionized diagnostic practices, ensuring greater accuracy, reproducibility and standardization5–7. Meanwhile, high-throughput multi-omics data, such as genomics, transcriptomics and proteomics, are accelerating our understanding of molecular func-tions in various physiological and pathological conditions8,9 and driving advancements in disease research and drug development10,11.
Analysing these large-scale biomedical data to drive scientific discoveries requires complex computational methods that integrate skills from bioinformatics, artificial intelligence (AI), software pro- gramming, statistics and mathematics12. These analyses are crucial for deciphering intricate genetic and molecular interactions, revealing biological mechanisms and disease origins13–15. Advanced machine learning and deep learning technologies further enable the discovery of novel biomarkers. AI-driven image interpretation, treatment response prediction and drug design highlight the role of computational tech-nologies in transforming biomedical data into actionable knowledge and enhancing clinical decision-making16–19.
To support these analytical needs, numerous computational tools have been developed for specific biomedical data tasks12,20,21. When integrated, these tools form complex workflows capable of processing diverse data types and producing notable results12,22. Platforms like Galaxy23, Nextflow22, Seven Bridges24 and DNAnexus25 aim to simplify workflow development and optimize the usage of these tools12. They make it easier for noncomputational users to assemble and execute workflows through user-friendly actions like clicking and dragging. However, these platforms still pose challenges, including dependency on predefined workflows that may reduce flexibility, outputs restricted to data files without interpretable summaries, and a lack of support for natural-language interaction, which restricts intuitive engagement. Large language models (LLMs) represent a transformative step towards general AI by substantially enhancing natural-language understanding26,27. LLMs have shown remarkable potential in inter-preting and reasoning, approaching human-level comprehension28–31.
This has led to the development of autonomous agents capable of understanding their environment, making decisions and taking actions accordingly32. Rapid progress in LLM-based agents has enabled them to understand and generate human-like instructions, profoundly shap-ing scientific research33, inspiring the concept of‘AI scientists’34. In the field of autonomous chemical studies, LLM agents have shown impressive performance for semi-autonomous experimental design and execution35, as well as powering autonomous mobile robots for exploratory synthetic chemistry.
The language capabilities and autonomous analysis abilities of LLM agents make them promising solutions to overcome the limita-tions of existing workflow platforms. For example, a GPT-4-powered agent was used to annotate cell types in single-cell RNA sequencing (scRNA-seq) analysis37. BioChatter was designed to develop custom bio-medical research software following open science principles38. Chat-GPT Advanced Data Analysis (ADA) streamlined automated machine learning for clinical studies39. AutoBA, an AI agent, was designed for fully automated multi-omic analyses40. CellAgent, a multi-agent framework powered by LLMs, automated processing and execution of scRNA-seq data analysis tasks41 . GenoTEX offered annotated code and results to solve gene identification problems42 . These innovative applications of LLM agents highlight their potential to assist in bio-medical data analysis.
Despite these advancements, current LLM agents for biomedical data analysis are constrained to specific tasks and data types, limiting their broader application. They struggle to effectively use diverse bioinformatics tools or chain them into workflows, making it chal-lenging to plan and execute complex, multistep analytical problems. Furthermore, they lack the ability to efficiently reuse past experi-ences. Evaluating these agents remains challenging due to the lack of well-established benchmarks, with BixBench43 being one of the few recently available. These limitations underscore the need for a more generalizable framework that can flexibly perform diverse biomedical data analyses and be systematically evaluated.
In this Article, we introduce BioMedAgent, a self-evolving LLM multi-agent framework, which learns to use various bioinformatics tools and chain them into executable workflows for carrying out diverse biomedical data tasks initiated by natural-language prompts. Given data and a scientific hypothesis, BioMedAgent functions as a virtual data scientist, performing multistep planning, autonomously explor- ing execution pathsand summarizing final conclusions. Its tool-aware flexibility, multi-agent interactive explorations (IEs) and self-evolving capacity enable robust performance across diverse biomedical data tasks when systematically evaluated on our newly constructed bench- mark and an external benchmark.
Results
LLM-powered multi-agent framework
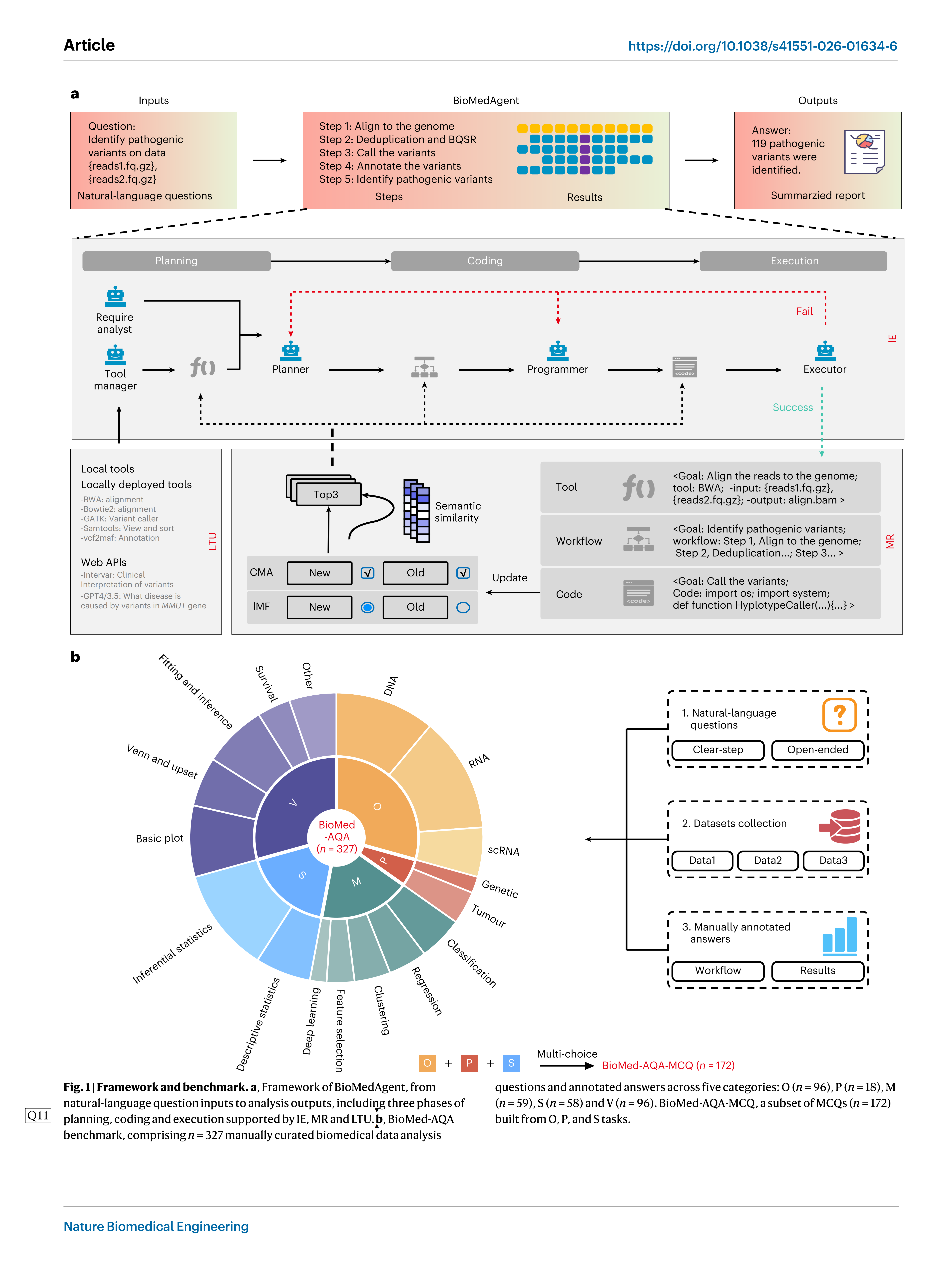
BioMedAgent is a multi-agent collaborative framework that uses LLMs such as GPT, DeepSeek and Gemini to generate diverse agents. These agents engage in IE through a three-phase collaborative process: planning, coding and execution (Fig. 1a). During the planning phase, multiple agents collaborate to interpret user requirements, select appropriate tools and design an effective analysis workflow. In the coding phase, agents generate the necessary code to implement each step of the workflow, using local tools or web application programming interfaces (APIs). The execution phase runs these codes in each step and produces results.
To enable self-evolving capabilities, BioMedAgent incorporates the memory retrieval (MR) algorithm, which uses a retrieval-augmented mechanism to facilitate there use of successful experiences. It system- atically records effective tool selections, workflows and executed codes, reusing and updating these memories with new insights as the system encounters similar tasks (Fig. 1a). This iterative cycle enables BioMedAgent to dynamically adapt and evolve overtime, consistently enhancing its problem-solving capabilities.
BioMedAgent supports natural-language input, enabling users to interact with the system using simple text instructions rather than structured commands. For example, a user can simply input:‘Identify pathogenic variantson data <reads1.fq.gz>, <read2.fq.gz>.’BioMedA-gent autonomously interprets the input and designs and executes the corresponding analysis workflows (Fig. 1a). This enables biomedical experts with no specialized computational or bioinformatics training to participate directly in their data analysis and interpretation, through natural-language interaction.
BioMedAgent’s strong performance across diverse tasks relies on its ability to interpret user instructions and activate the appropriate tools. This is accomplished by the Tool Manager agent, which man-ages both locally deployed tools and web APIs within its workspace (Fig. 1a). These tools are crucial for specialized bioinformatics analyses, particularly inomics and precision medicine44. While the Programmer agent, derived from foundational LLM models, possesses general coding ability, it struggles to generate de novo implementations of specialized tools45. The Tool Manager agent addresses this by enabling LLMs to learn how to use existing tools from documentation covering descriptions, inputs, outputs and dependencies rather than rewriting new codes. Once integrated, these tools become available for autono-mous use during task execution.
BioMed-AQA: benchmark of biomedical data tasks
To comprehensively evaluate the performance of BioMedAgent, we developed BioMed-AQA, a benchmark that includes 327 biomedical data analyses questions across five categories: omics analysis (O), pre-cision medicine support analysis (P), machine learning (M), statistical analysis (S) and data visualization(V) (Fig. 1b). This benchmark is based on 17 types of biomedical data analysis tasks, including DNA, RNA, scRNA, classification, regression, deep learning and others. These diverse task types were selected to comprehensively assess BioMedA-gent’s performance across a wide range of biomedical analysis contexts.
We categorized the instructive questions in BioMed-AQA into clear-step and open-step types to test BioMedAgent’s ability to trans-late natural-language inputs into actionable steps. Clear-step questions are those with well-defined steps (for example,‘Align sequencing data <tumor.R1.fastq.gz, tumor.R2.fastq.gz, normal.R1.fastq.gz, normal. R2.fastq.gz> using BWA, call somatic mutations with GATK’s Mutect2, annotate variants with vcf2maf, and recommend therapy based on the genomic variants.’). Open-step questions are those in which only the objective is introduced, and the tools or steps are not clearly specified (for example,‘Perform mutation analysis on sequencing data <tumor. R1.fastq.gz, tumor.R2.fastq.gz, normal.R1.fastq.gz, normal.R2.fastq.gz> from cancer patients and recommend therapy.’).

Combined manual and automated evaluation using Win score
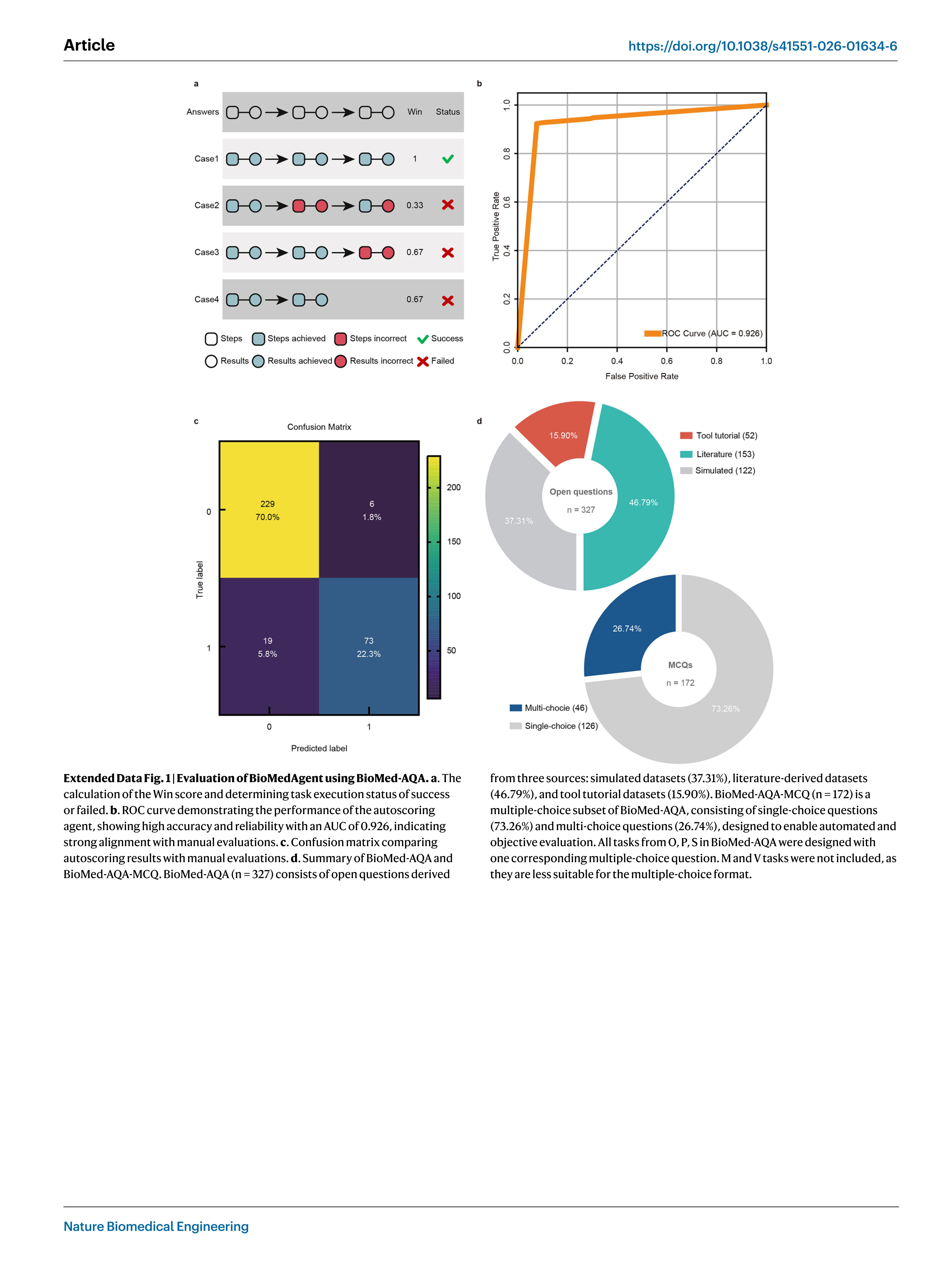
The effectiveness of BioMedAgent is assessed using a‘Win’score, designed to measure the system’s ability to achieve predefined mile-stones (Extended Data Fig. 1a). Each of the 327 biomedical analysis questions is comprehensively annotated with the necessary steps and corresponding milestones for successful execution. A task that success-fully meets all designated milestones receives a‘Win’score of 1, while a task that meets only one out of four milestones is scored as 0.25. A‘Win’score of 1 indicates an execution status of‘success’, whereas a score below 1 is considered an execution status of‘failed’. The overall success rate is determined by calculating the percentage of tasks that receive a‘Win’score of 1.
Evaluating agentic AI is still a major challenge, especially for open-ended biomedical data tasks where there can be multiple cor-rect ways to solve a problem. As observed in previous works such as AutoBA40, BIA46, BioMaster47 and CellAgent41, manual evaluation remains the most reliable approach, despite being labour intensive and limited in objectivity. In BioMed-AQA, manual assessment serves as the primary method, using the Win score above. This approach is neces-sary because the inherent uncertainty of generated content by agents often makes exact matching impractical—for example, differences in decimal precision (for example,‘0.545’versus‘0.55’), formatting issues (for example,‘1/3’versus‘1:3’) or different visual representations (for example, survival curves created using different packages). Manual evaluation effectively addresses these challenges by considering con- textual correctness rather than rigid exact matching.
To improve objectivity and scalability, we also introduced an autoscoring agent that compares the outputs against predefined milestones. It showed 92.3% consistency with manual evaluations (AUC = 0.926, P < 0.001; Extended Data Fig. 1b,c), reducing reliance on manual review. Furthermore, we also developed BioMed-AQA-MCQ, a 172-question multiple-choice questions (MCQs) subset, each con-taining five options, that supports fully automated, unambigu-ous evaluation (Fig. 1b and Extended Data Fig. 1d). Task from O, P and S categories in BioMed-AQA were supplemented with MCQs, while M and V tasks were not included, as they are less suitable for MCQ format.
Enhanced overall performance on O, P, M, Sand V tasks
BioMedAgent has effectively shown its capability in handling general biomedical data analysis tasks, achieving an impressive overall aver-age success rate of 77% across various categories on the BioMed-AQA benchmark (Fig. 2a). This includes high success rates in specific cat-egories: 94% inO, 78% in P, 90% in M, 59% inS, and 65% in V (Fig. 2a,b).
We compared BioMedAgent’s performance with other well-known LLM agents, all using the same underlying model, GPT-4omini. BioMed-Agent achieved a superior success rate, with consistent improvements across O, P, M, Sand V tasks, outperforming two of OpenAI’s web agents and one local-tool-based agent. By contrast, the online ChatGPT-4omini was unable to provide data analysis, while the GPT Assistants achieved a 39% success rate, and GPT Function Call yielded a 33% success rate (Fig. 2a). Remarkably, it also outperformed the higher version model, ChatGPT-4o, which achieved a success rate of 46%, showing consistent improvements in the O, P, M and V tasks. By using local tools within its workspace, BioMedAgent showed a notably broader analytical scope compared to online ChatGPT-4o and GPT Assistants, achieving a 100% analysable scope. This exceeded the 68% and 65% analysable scopes observed for the latter two (Fig. 2a).
When tested with different open LLMs, including DeepSeek v3 and Qwen3, BioMedAgent maintained robustness, achieving 77% and 75% accuracy, respectively (Fig. 2a). The accuracy on MCQs was 76%, comparable to that on open questions (Fig. 2a). Although MCQs require completing the analysis before answering, they allow for some correct guesses, which explains the similar performance compared to open questions.
BioMedAgent’s overall high performance remains consistent even as the complexity of workflows increases. Specifically, for P tasks, which typically involve an average of 6 planned steps, BioMedAgent continues to maintain a high success rate (Fig. 2a–c). While showing robust performance overall, BioMedAgent exhibitedits lowest perfor-mance on S tasks. Although it surpassed other agent applications using ChatGPT-4omini, its success rate was lower than that of ChatGPT-4o, which uses alarger LLM model. This relatively weaker performance of BioMedAgent for S tasks maybe attributed to the inherent demands of statistical tasks, which require fundamental data or table processing capabilities and thus rely more on the underlying LLMs.
Local tool usage significantly enhances the success rate
For biomedical data analysis tasks such as O and P, the use of specialized bioinformatics tools is crucial for conducting analysis effectively44,48. However, these tools often require complex environments and cannot be easily executed using standard Python packages, making it difficult for LLMs to program them correctly. To address this challenge, we enhanced BioMedAgent by establishing a Tool Manager agent respon-sible for learning and maintaining specialized tools with its workspace. This process involves integration of specific tools, which requires learn-ing their documentation, including descriptions, inputs and outputs and preparing the necessary dependencies for execution. With this knowledge, BioMedAgent can activate the appropriate tools from the Tool Manager as needed.
We have integrated 67 specialized bioinformatics tools into BioMedAgent, mainly in the O, P and M categories. This integration has significantly improved BioMedAgent’s performance, with average success rates for O, P and M tasks exceeding 90% (Fig. 2a,b). Moreo-ver, tasks with local tool usage (LTU) show significantly higher Win scores compared to those without LTU across all tasks (P= 1.259 × 10_4, two-tailed t-test) and M tasks (P = 1.477 × 10_3, two-tailed t-test) (Fig. 2d and Extended Data Fig. 2a), with a similar trend observed in the success rate (Extended Data Fig. 2b). In cases where a required tool is missing, the agent can generate a custom tool code (CTC) to fill the gap. LTU and CTC complement eachother, with LTU used exclusively in 46.25% of successful tasks, CTC in 28.85% and a combination of both in 24.90% (Extended Data Fig. 2c).
High robustness to varied prompts from human medical experts
Different users often phrase the same analytical tasks in their unique ways, influenced by their individual backgrounds and preferences. To evaluate how well BioMedAgent can handle these semantic vari- ations, we conducted a test involving three human medical experts. Each human expert rephrased existing benchmark questions in more natural and varied ways without changing their analytical objectives. This allowed us to assess how well BioMedAgent understands the same task when expressed differently. The accuracy obtained from the three experts (0.761, 0.752 and 0.758) were comparable to the original bench-mark accuracy (0.774) (Fig. 2e), indicating that BioMedAgentis robust to linguistic variations.
We further examined whether BioMedAgent can translate broad instructions into actionable analytical processes, by comparing its performance on clear-step and open-step questions that describe the same task with different levels of detail. Comparable Win scores were observed between the two types of questions across the total (P = 0.622, two-tailed t-test; Fig. 2f) and O, P, M, S and V tasks (Extended Data Fig. 2d); the success rates showed the same pattern (Extended Data Fig. 2e). The results suggested BioMedAgent’s robust capability to autonomously plan and execute necessary processes, even in the absence of explicit step-by-step guidance. This adapt-ability highlights the system’s advanced understanding, making it suitable for users without specialized computational or bioinformat-ics knowledge.

Multi-agent collaboration enhanced by the IE algorithm
Human group intelligence involves collaborative explorations by mul-tiple individuals to handle complex problems while using existing experiences to address similar problems49. BioMedAgent enhances the planning and execution of tasks by emulating human group col-laboration, thereby increasing problem awareness and facilitating appropriate actions. While encountering problems for the first time, BioMedAgent activates IE to expand the explorations and optimize task planning and execution. While encountering problems again, it trig-gers MR to use the past experiences. In this way, IE uncovers potential solutions, while MR reduces redundant explorations (Fig. 1a).
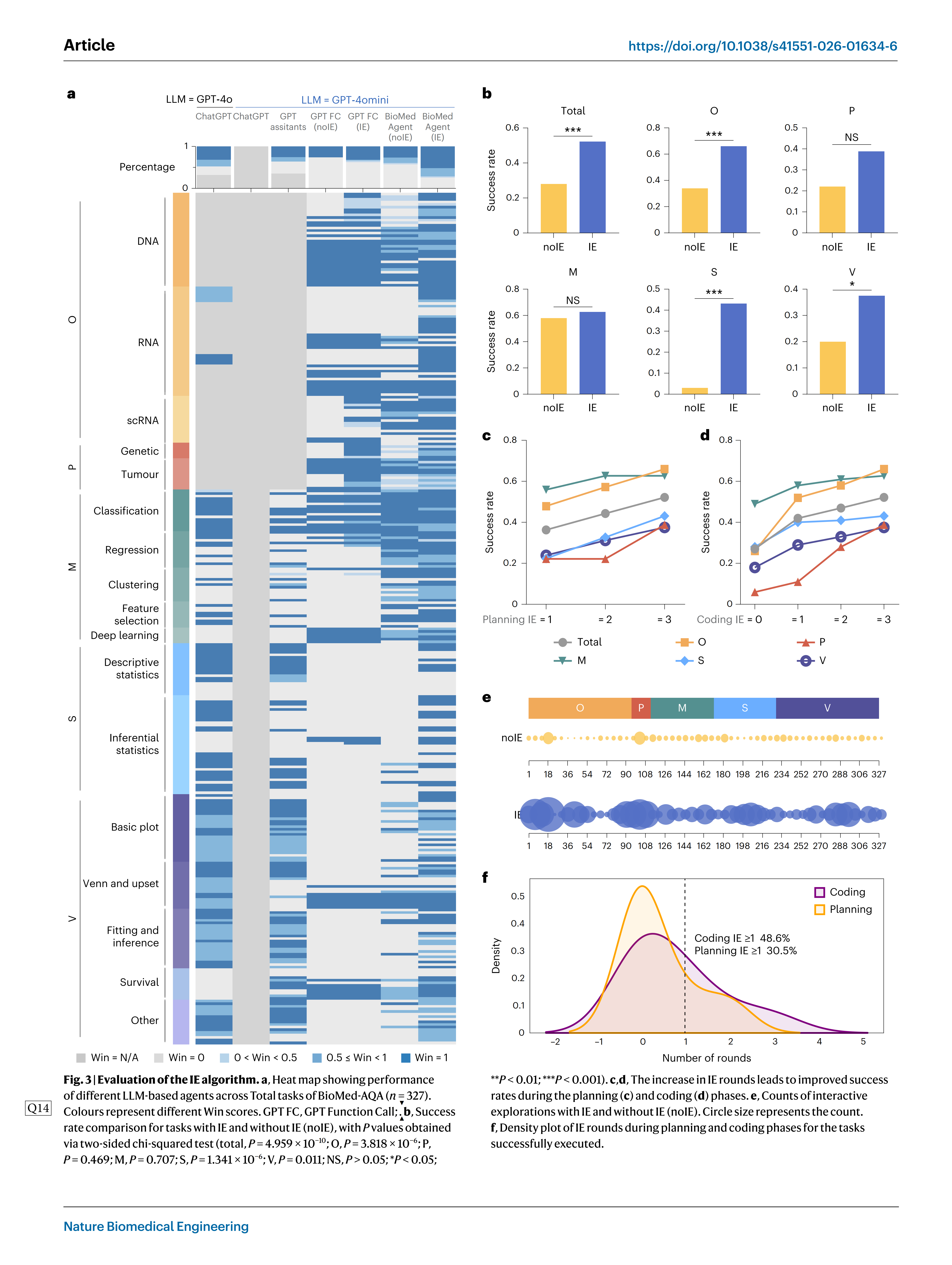
BioMedAgent uses multi-agent collaboration enhanced by the IE algorithm to effectively address biomedical data analysis challenges (Extended Data Fig. 3a). This method has significantly improved the Win score, allowing more tasks to be analysed correctly. BioMedAgent with IE achieved the highest average Win score, outperforming noIE (without IE) strategy and surpassing two of OpenAI’s web agents, as well as one local agent (Fig. 3a and Extended Data Fig. 3b,c). Theaver-age success rate nearly doubled, increasing from 28% to 52% when compared to noIE strategy (P = 4.959 × 10_10, two-sided chi-squared test; Fig. 3b). Consistent improvements were observed across five categories of tasks: O increased from 34% to 66% (P= 3.818 × 10_6, two-sided chi-squared test), P from 22% to 39% (P= 0.469), M from 58% to 63% (P= 0.707), S from 3% to 43% (P= 1.341×10_6) and V from 20% to 38% (P= 0.011) (Fig. 3band Extended Data Fig. 3d).
As the number of IE rounds increased in both the planning and coding phases, the success rates consistently improved (Fig. 3c,d). IE expands the potential solution space and promotes innovations in group intelligence systems49. With the IE algorithm, the total number of explorations increased from 916 to 2,696, almost tripling, indicating that a significantly greater number of potential solutions were auto- matically explored (P = 7.468×10_15, two-tailed paired t-test) (Fig. 3e). Over 30.5% of successful tasks required extra interactions (rounds of IE ≥ 1) during the planning phase, while 48.6% involved extra interac-tions (roundsofIE ≥ 1) during the coding phase (Fig. 3f). This collabora-tive strategy enabled approximately 60.8% of previously failed tasks to succeed, highlighting the IE algorithm’s effectiveness in boosting BioMedAgent’s performance.
Self-evolving through the MR algorithm
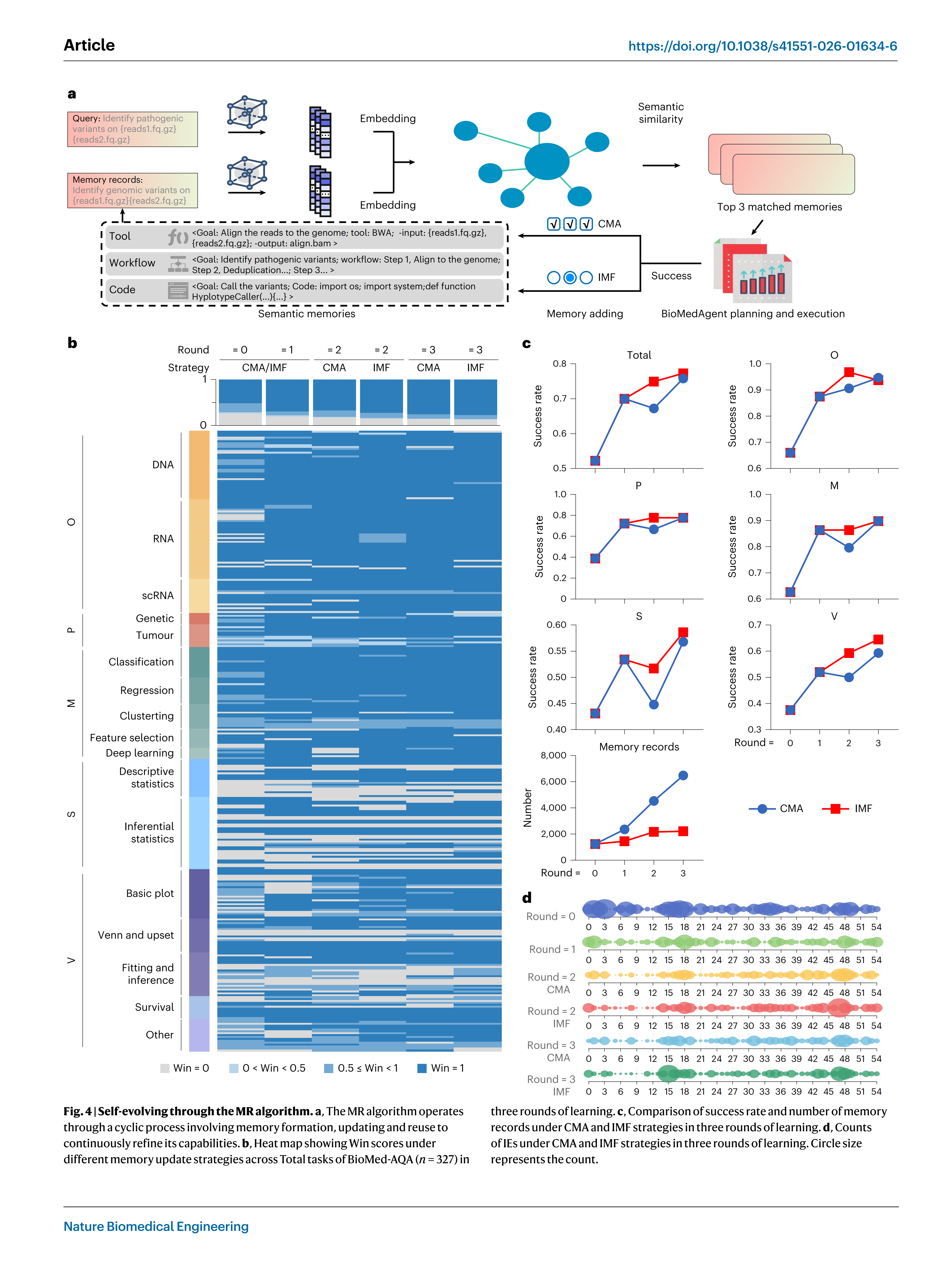
The MR algorithm enables BioMedAgent to be self-evolving by lev-eraging a dynamic memory base built from previous tasks, allow-ing past experiences to be reapplied in new contexts (Fig. 4a and Extended Data Fig. 4a). When evaluated over three rounds of iterative learning using the BioMed-AQA benchmark, BioMedAgent showed pro-gressive performance gains (Fig. 4band Extended Data Fig. 4b), with the success rate rising from 52% to 77% (Fig. 4c). Across the five task catego-ries, the success rate increased from 66% to 94% for O (P= 3.103 × 10_6, two-sided chi-squared test), 39% to 78% for P (P= 0.043), 63% to 90% for M (P= 0.001), 43% to 59% for S (P= 0.137) and 38% to 65% for V (P= 3.070 × 10_4) (Fig. 4c). BioMedAgent retrieved relevant past expe-riences through semantic similarity queries, in which analyses of the same type show closer semantic relationships, facilitating their appli-cation to related tasks (Extended Data Fig. 4c).
To ensure that the self-evolving did not result from direct memory access to identical questions and the subsequent recall of the answers, we further randomly reserved 30% of the BioMed-AQA benchmark as unseen questions and used the remaining 70% as seen questions. We performed iterative MR-based learning using only the seen subset and subsequently evaluated its performance on unseen subsets to see the system’s generalization. The success rate showed a slight decrease for unseen questions compared to seen questions (69% versus 76%, two-sided chi-squared test, P = 0.307) yet remained significantly higher than before learning (69% versus 52%, two-sided chi-squared test, P = 0.019) (Extended Data Fig. 4d). These results confirm that the MR supports generalizable self-evolving, improving the agent’s capabili-ties in toolselection, planning and coding, rather than relying on direct recall of previously generated answers.
Effective memory update strategies enable memory records to be optimally queried and applied indifferent analytical contexts50. BioMed-Agent has established two distinct memory update strategies: continu-ous memory accumulation (CMA), which retains all successful records, and iterative memory forgetting (IMF), which selectively prunes out-dated records (Fig. 4a and Extended Data Fig. 4a). IMF achieved faster convergence and slightly superior final success rates compared to CMA across all task categories (Fig. 4c and Extended Data Fig. 4e). It also showed greater stability in iterative rounds of learning, whereas CMA showed an abnormal decline in performance between the first and second rounds (Fig. 4c and Extended Data Fig. 4e). Furthermore, IMF required less intermediate memory, making it more economical in the long-term operation and maintenance of an expanding platform (Fig. 4c). Both IMF and CMA memory update approaches could reduce redundant re-explorations (P= 1.646 × 10_7 between rounds 3 and 0 for IMF, P= 6.759 × 10_8 for CMA, two-tailed paired t-test) (Fig. 4d).
After three rounds of learning with IMF, BioMedAgent achieved high performance across BioMed-AQA (Fig. 2a,b and Extended Data Figs. 5–8), showing its self-evolving capability for continuous improvement.
Robust generalization on external benchmark
Evaluating agentic AI is challenging, especially for biomedical data tasks where the same problem can be solved in several valid ways and the outputs cannot be judged by exact matching. There are few well-established benchmarks designed specifically for assessing sci-entific agentic AI. BixBench43 is one of the few publicly available bench-marks, offering a comprehensive dataset over 50 real-world analytical scenarios (‘capsules’) and 296 associated questions. Its scale is compa-rable to BioMed-AQA, and it coversawiderange of bioinformatics tasks.
To evaluate BioMedAgent’s generalizability beyond our curated BioMed-AQA benchmark, we tested it on BixBench, including both open questions and MCQs. When running without domain-specific tools and relying solely on CTC, BioMedAgent achieved 49% accuracy on open questions, 49% on MCQs with refusal and 63% on MCQs without refusal. These results significantly outperformed the benchmark-provided agent baselines of 37%, 42% and 55%, respectively (P = 5.459 × 10_5, 2.119 × 10_2, 2.814 × 10_2, two-sided McNemar’s test) (Fig. 5a). These find-ings show BioMedAgent’s strong capability in autonomous multistep biomedical analysis, even without tool integration, and highlight its robust generalization to external benchmarks.
Comparing with existing multi-omicsAI agents
Several multi-omicsAI agents have been recently developed, including BioChatter38, BixBench43, AutoBA40, BIA46, BioMaster47, OLAF51, ADA39, GenoTEX42, CellAgent41 and CASSIA52, each designed to solve specific biomedical data analytical tasks. To better illustrate the strengths of BioMedAgent, we conducted a structured comparison with these 10 existing multi-omicsAI agents across three dimensions: functionality, extensibility and evaluation. Functionality is defined by the core capa-bilities such as planning, coding, execution and natural-language inter-action. Extensibility refers to the ability to manipulate tools, support inter-agent communication and evolve overtime. Evaluation focuses on the benchmark scale and assessment methods.
Most existing agents support the end-to-end process of planning, coding and execution but are often limited to specific analysis types (Fig. 5b). Among the more general agents, AutoBA provides support for omics analysis, while BioMedAgent shows broader applicability across O, P, M, S and V tasks. BioChatter also supports more general tasks by using knowledge graphs and live API services to enhance question-answering capabilities and to facilitate the development of biomedical software and databases within an open science framework.
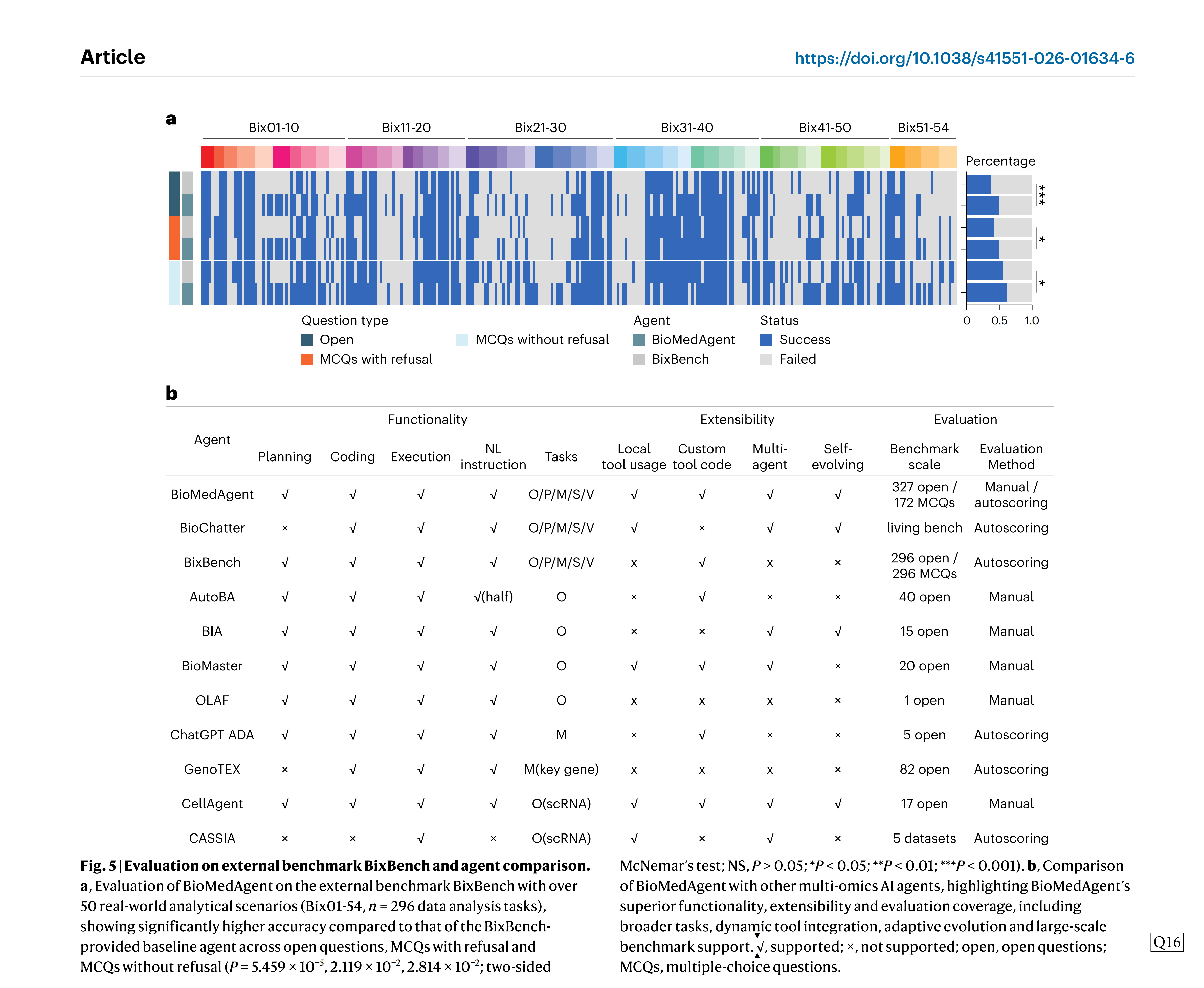
Extensibility remains limited in most agents, as few support dynamic tool integration and adaptive evolution. By contrast, BioMed-Agent combines both LTU and CTC, reinforced by a self-evolving mech- anism via the MR algorithm. Regarding evaluation, only BixBench and BioMedAgent offer large-scale benchmarks (300 tasks). Most others rely on small application demos. While open questions are the dominant evaluation method, only BixBench and BioMed-AQA include MCQs to support automated and objective evaluation. Moreover, most existing evaluations depend on manual annotation, whereas BixBench and BioMedAgent support autoscoring assessment.
In summary, BioMedAgent offers broader task coverage, greater flexibility in tool usage, adaptive evolution and amore systematic and scalable evaluation framework compared to existing agents.
Applying BioMedAgent to cross-omics analysis of RNA-seq and scRNA-seq data
To assess BioMedAgent’s ability to assist in complex biomedical research, we designed a cross-omics question: From which cell types are the highly differentially expressed genes (DEGs) in non-small cell lung cancer (NSCLC) derived? This requires integrating molecular and cellular information.
We downloaded bulk RNA-seq data from 67 NSCLC samples (34 tumour and 33 adjacent normal samples) and 22 scRNA-seq matrix files from the Gene Expression Omnibus (GEO, GSE268175, GSE131907). Using three natural-language instructions, BioMedAgent conducted large-scale cross-omics data analysis, covering bulk RNA-seq, scRNA-seq and cross-omics integrative mining. It automatically pro-grammed and executed the planned steps, which included trimming, alignment, quantification, visualization for RNA-seq, normalization, scale, clustering, marker identification for scRNA-seq and filtering, intersection and visualization for cross-omics analysis (Fig. 6a).
Atotal of 1,831 DEGs, including 1,309 upregulated and 522 down-regulated, were identified from bulk RNA-seq data, with corresponding volcano plot and heat map for visualization (Fig. 6b). These covered 78% (n = 842) of the DEGs identified by the GEO2R online tool, show-ing consistent findings (Extended Data Fig. 9a). Eight cell types were detected, and cell-specific DEGs were identified from the scRNA-seq data (Fig. 6b). Cross-omics integration revealed NSCLC-associated cell-specific DEGs:ABCC3, SERINC2 and SEZ6L2, which were predomi-nantly overexpressed in epithelial cells (Fig. 6b). ABCC3 encodes a protein belonging to the MRP subfamily which is involved in multi-drug resistance and plays a key role in NSCLC prognosis53. SERINC2 is associ-ated with cervical cancer54,55, and SEZ6L2 is upregulated in lung cancer, which can be used for predicting the prognosis56,57.
Using natural-language instructions, BioMedAgent autonomously conducted a large-scale cross-omics analysis and produced results

consistent with outputs from official online tools and with the pub-lished literatures.
Streamlined machine learning modelling for cancer-associated venous thromboembolism prediction
Machine learning is difficult for biomedical experts due to the complex requirements of data preprocessing, model selection and outcome evaluation. To test whether BioMedAgent can simplify this using natural language, we reproduced a Nature Medicine study58 that built a liquid biopsy model using circulating tumour DNA (ctDNA) to assess the risk of venous thromboembolism (VTE) in cancer patients.
Using natural-language instructions, BioMedAgent automatically generated and executed a complete end-to-end machine learning modelling workflow for this task, encompassing model training, pre- diction, performance evaluation and comparison. It reproduced the entire analytical process described in the publication, from hypothesis formulation to model validation and interpretation, without manual coding. The results validated the hypothesis and confirmed the con-clusions of the original study, showing that both the LB+ model (liquid biopsy plus two minimal clinical variables) and the All model (full model including all variables) showed superior predictive capabilities, with c-index values of 0.73 and 0.74, respectively (Fig. 6c). By contrast, the Khorana score model showed a lower predictive performance with a c-index of 0.61. These results showed that liquid biopsy-based models significantly outperform the Khorana score model in predict-ing VTE risk.
Overall, BioMedAgent enables biomedical researchers to run complex machine learning pipelines directly from natural language, bridging the gap between clinical expertise and computational technologies.
Improved cell segmentation and classification on pathological images
Cell segmentation and classification is crucial for pathological image analysis, as precise cell segmentation results can support downstream tasks such as lesion identification and survival assessments. However, cell segmentation tasks in pathology face challenge soflow-resolution images. Recent deep learning algorithms can enhance resolution by transforming low-resolution images into high-resolution. We used BioMedAgent to integrate cell segmentation with resolution enhance-ment algorithms, aiming to enhance the effectiveness of cellsegmen-tation. We guided the construction of an automated workflow by incorporating MiHATP v.1.059, a resolution enhancement tool, into BioMedAgent’s local workspace. BioMedAgent’s planned workflow automated data splitting, image resolution enhancement, cell seg-mentation and Dice evaluation (Extended Data Fig. 9b).
Compared to the baseline model, BioMedAgent showed consistent improvements in absolute Dice scores across four cell types: inflamma- tory, epithelial, spindle-shaped and others, with an overall Dice gain of +0.86% (Fig. 6d). In addition to the absolute gain, we also calculated the relative improvement compared to the remaining gap to the upper limit (referred to as‘Delta Dice to upper’). The results showed that the remaining gap closed reached 29.9% overall. These absolute and relative gains showed the practical value of integrating BioMedAgent with resolution enhancement algorithms, consistently improving segmentation accuracy across various cell types.
Local deployment and intuitive chat interface
BioMedAgent can run locally while calling online LLMs (GPT-4/4o/4omini, Claude 3) or local open-source models (for example, Qwen3-235b-a22b). It provides an intuitive chat interface that makes it easy to start tasks, monitor execution and view results, so complex workflows become manageableona local set-up (Extended Data Fig. 10). This design is lightweight, easy to deploy, and scalable, making BioMed-Agent usable for researchers with diverse technical backgrounds.
Discussion
BioMedAgent represents an important step toward autonomous, data-centric biomedical analysis by using a flexible multi-agent sys-tem architecture. Unlike conventional LLM-based approaches that primarily generate textual responses or isolated code fragments, BioMedAgent reconfigures LLMs into specialized, interacting agents capable of end-to-end data workflows. Rather than functioning as a question-answering system, it autonomously plans, codes, executes and adapts, thereby bridging the gap between AI automation and prac-tical biomedical data research needs.
A core advantage of BioMedAgent lies initstool-aware flexibility. Unlike general-purpose multi-agent frameworks such as LangGraph, CrewAI and SmolAgents, which rely on fixed agent roles and predefined tools, BioMedAgent supports dynamic integration of both LTU and CTC. By learning tool usage directly from documentation and adapt-ing to task-specific requirements, the agent can incorporate validated domain-specific tools when available or generate custom code to address new or unsupported analytical steps. This dual mechanism ensures high robustness in established scenarios and strong adapt-ability when faced with novel tasks.
BioMedAgent’s intelligence arises from its multi-agent and self-evolving design. Through the IE algorithm, agents iteratively exchange feedback and context across planning, coding and execu-tion phases, enabling rapid, targeted explorations and adjustments. Each agent maintains a dedicated memory space, storing successful experiences and using the MR algorithm to recall and reuse the past experience. This continuous learning process enables BioMedAgent to improve autonomously overtime.
An important design is the balance between exploration and expe-rience reuse. Exploration can lead to new discoveries but consumes resources, while relying on past experience may limit innovation49. BioMedAgent currently adoptsa stratified approach based on semantic similarity, automatically determining whether to try new explorations or use existing experiences, without manual tuning. In future iterations, introducing a user-controllable index could allow dynamic adjustment of this balance, providing finer control over computational cost and innovation.
Another consideration is the integration of both positive and nega-tive experiences. Currently, BioMedAgent, like most LLM-based sys-tems, preferentially records successful entries60. However, failure cases provide valuable insights for avoiding repeated mistakes and enhancing robustness50. Understanding and integrating these unsuccessful experi-ences is also meaningful, and future research aimed at more effective use of‘negative experiences’could be beneficial for BioMedAgent61.
The efficiency of memory update mechanisms is critical for sus-taining long-term self-evolving62–64 . Our findings indicate that the IMF strategy yields faster and more effective adaptation than CMA, although the results are based on limited rounds and task sets. More advanced memory update strategies, such as mathematical scoring, could be further investigated as the system evolves and expands.
BioMedAgent still has limitations that are common in LLM-based systems, particularly hallucinations arising from next-token probabil-istic modelling. In multistep workflows, it may fabricate intermediate results or final conclusions by failing to correctly track previously generated outputs. In addition, the performance of the agent system is inherently limited by the reliability of the specialized tools within its workspace, and scaling up tool numbers may introduce computational bottlenecks. Future work should focus on reducing hallucinations to enable trustworthy reasoning, developing a more standardized tool infrastructure and evolving from closed-loop digital interactions to more open real-world engagement.
In summary, BioMedAgent advances the vision of an ‘AI data sci-entist’, capable of autonomously conducting complex, data-centric biomedical research and continually improving through accumulated experience.
Methods
BioMed-AQA benchmark
The datasets in BioMed-AQA were manually curated from three sources:
(1) simulated datasets, (2) tutorial examples from commonly used bioinformatics tools or packages and (3) literature-derived datasets from biomedical research. Among them, simulated datasets account for 37.3%, tool tutorial datasets for 15.9% and literature-derived data-sets for 46.8%. Together, these datasets cover five categories of O, P, M, Sand V tasks.
O and P task datasets. In the DNA-sequencing (DNA-seq) analysis,we used VarBen v.1.0 (ref. 65) to simulate DNA-seq data that include both germline and somatic tumour mutations. First, we downloaded whole exome sequencing data from the National Institute of Stand-ards and Technology Genome in a Bottle consortium (NA12878)66 and aligned it to the genome using the GATK v.4.0 Best Practices workflow to obtain alignment files in Binary Alignment Map format. We then used VarBento feature pathogenic germline mutations on MMUT, PAH and SLC22A5 genes and oncogenic somatic tumour muta- tions onEGFR, KRAS and BRAF genes, resulting in three germline and three tumour sequencing datasets, together with control samples. Precision medicine datasets were generated from these DNA-seq data to support genetic disease prediction and drug analysis in tumour samples.
For RNA-seq, we used three sets of expression profile data: one microarray dataset from the GEO database (GSE19089) and two next-generation sequencing datasets from the University of Califor-nia, Santa Cruz Xena Browser (https://xenabrowser.net/), including The Cancer Genome Atlas Colon Adenocarcinoma and The Cancer Genome Atlas Rectum Adenocarcinoma. Cancer and normal control labels from the raw data were used as grouping information.
For scRNA-seq, we used three datasets: the first datasetisa periph-eralblood mononuclear cell (PBMC) dataset provided by 10X Genomics and used in the Seurat tutorial67, containing approximately 2,700 single cells sequenced on the Illumina NextSeq 500; the second dataset is oesophageal cancer data obtained from a published study68; the third dataset is another PBMC dataset provided by 10X Genomics, used in the Scanpy tutorial69, containing approximately 68,000 single cells sequenced on the Illumina NextSeq 500.
M task datasets. Machine learning includes classification, regression and clustering tasks. For classification, three datasets from Kaggle (https://www.kaggle.com/datasets) were used: the Diabetes dataset, the Heart Attack Risk Prediction dataset, and the Predicting Heart Fail-ure dataset. For clustering, the Breast Tissue dataset from the UC Irvine Machine Learning Repository (https://archive.ics.uci.edu/datasets) and the two Kaggle datasets (the Heart Attack Risk Prediction dataset and the Predicting Heart Failure dataset) were used, with classification labels removed for clustering. For regression, three datasets from Kag-gle—the Medical Cost Personal Datasets, the Diabetes dataset and the Heart Attack Risk Prediction dataset—were used to predict medical expenses and the risk of disease occurrence.
S and V task datasets. For statistical analysis, the t-test task datasets were obtained from the demos in the R v.4.3.0 package for‘stats::t. test’. The Wilcoxon-test task datasets are from three built-inR datasets: airquality, birthwt and iris. The LEfSetask datasets were derived from our previously published microbiome data. The survival task datasets were from TCGA, including ovarian carcinoma, liver hepatocellular carcinoma and lung adenocarcinoma and include curated phenotype information such as stage, gender and KRAS/EGFR/ALK mutation status. For data visualization, the enrichment bubble plot task dataset was from the demos provided on the KOBASv.3.0 website70, and the Venn plot task datasets were from the official examples provided in the R package Venn Diagram.
Evaluation design
Open questions and MCQs. The BioMed-AQA benchmark contains both open questions and MCQs to comprehensively evaluate BioMedA-gent’s performance in biomedical data analysis. The open questions (n = 327) are designed to reflect real-world bioinformatics scenarios, with annotated milestones provided as reference answers. By con-trast, the BioMed-AQA-MCQ (n = 172) are more structured and repre-sent a subset of BioMed-AQA, as M and V tasks are excluded because they are not suitable for a MCQ format. This subset includes 46 MCQs and 126 single-choice questions, each offering five answer options (Extended Data Fig. 1d). During testing, we did not indicate whether the question was single-choice or multiple-choice to minimize the likelihood of chance-correct responses.
Comparison against OpenAI agents. We conducted a comparative evaluation with three OpenAI agent frameworks: ChatGPT, Assistants API and Function Calling (using GPT-4o and GPT-4omini models). As OpenAI’s underlying agent framework is closed-source, we relied on their official online services or API for this comparison. ChatGPT func-tions as OpenAI’s public-facing interactive interface; we submitted tasks along with the input data through the chat window. Assistants API is part of OpenAI’s hosted agent service, which supports file uploads to the user workspace; we used the API to upload input data, send analysis prompts in batches, and write and execute Python code through its Code Interpreter to iteratively handle complex analyses. Function Calling allows for invoking external tools by creating local APIs and exposing them to the agent through structured definitions; we tested its ability by using local tools (same as we used in BioMedAgents) provided in its environment. We used the identical task promptsand data inputs from BioMed-AQA to ensure consistency in comparison with BioMedAgent.
Evaluation on external benchmark BixBench. We tested BioMedA-gent’s performance on the external benchmark BixBench, which is designed to assess agentic AI in bioinformatics tasks within a controlled Jupyter Notebook environment. We followed BixBench’s official pro- tocolby using its built-in agent, which operates through three primary commands: edit cell (to modify and execute code cells), list workdir (to inspect the workspace directory structure) and submit answer (to submit final responses). We configured BioMedAgent with GPT-4.1 as the base LLM and integrated the required data files into the Jupyter Notebook environment. The original BixBench questions were used as prompts to initiate data exploration and answer generation. The evaluation was conducted in three parallel testing groups: 296 open questions, 296 MCQs with refusal and 296 MCQs without refusal.
Tool-aware agent framework
Learning to use domain-specific tools. BioMedAgent allows users to add specialized tools into its localized workspace, thereby enabling effective use for various tasks. By learning tool documentation and preparing dependencies, tools can be integrated into the BioMedA-gent’s local workspace.
Tool documentation. The documentation includes six parts: name, identifier, description, input, output and example. The name specifies the tool’s title. The tool identifier corresponds to the function name in the code and must remain consistent so that the LLM can be aware of the tool’s presence within its workspace. The description providesan overview of the tool’s functionality. The input and output cover details such as parameter names, types and descriptions. In addition, each documentation includes a simple contextual example of how to invoke the tool, helping BioMedAgent better understand its proper usage.
Containerization and central API. To manage the diverse implementa-tion languages and environmental dependencies of different tools,Docker containerization technology is used to create isolated execu-tion environments for each tool. Python is then used to create a central API that acts as the main controller for initiating the program. This API handles input processing, launching the container, executing the com-mand and retrieving results from a predefined workspace. This design ensures consistent, reproducible execution across heterogeneous tool environments.
Two-round tool retrieval via Tool Manager agent. A user’s initial task request typically includes the user’s available resources and their expected goals G, but often lacks specific details about the analytical process. Fora given local tools set T = {t1, t2, t3} in BioMedAgent’s work- space, the Tool Manager evaluates the relevance of each tool using the user’s initial description P and the tool documentation D = {d1, d2, d3}. This process generates a relevance score ranging from 0 to 1, reflect-ing how suitable each tool is, and forms the initial relevance score set S = {s1, s2, s3}. Tools with high relevance scores are selected for further exploration and usage recommendation.
To balance tool retrieval efficiency with context usage, we allowed users to specify acut-off on the initial relevance scores. A higher cut-off improves tool retrieval precision but narrows recall and reduces the token cost of subsequent reasoning rounds (Supplementary Fig. 1a). Using 184 BioMed-AQA tasks that were successfully executed with LTU, we treated the tools actually invoked in these tasks as reference tools and evaluated different cut-offs for their ability to retain needed tools while limiting token cost. A cut-off of 0.6 provided the most favour-able balance between recalling reference tools and controlling token cost (Supplementary Fig. 1b). In practice, this cut-off can be flexibly adjusted within 0.4–0.8 according to workspace size and available context budget.
The selected high-scoring tools are then used to generate usage recommendations, including relevant documentation and their upstream and downstream dependencies. These recommendations act as workflow‘seeds’guiding subsequent workflow expansion. Based on these results, the Tool Manager performs a second round of rel-evance assessment torefine the tool list. In this step, outlier tools with excessively high scores may be removed, while initially low-scoring but essential intermediate tools can be reintroduced by‘seeds’guid-ing. This two-round strategy helps BioMedAgent overcome common LLM limitations related to token capacity and long-context reasoning, ensuring that the final workflow includes the most necessary tools (Supplementary Fig. 2a).
Combining LTU and CTC. Compared to existing multi-omics agents like autoBA and general-purpose frameworks like LangGraph, which are often constrained by limited predefined tool usage, BioMedA- gent provides much greater flexibility by combining LTU and CTC (Supplementary Fig. 2b). It integrates tools into the workspace on demand by learning their documentation and dependencies, allowing the agent to dynamically bind and invoke tools at runtime without rely-ing on fixed configurations. When a required tool is missing, the agent can generate CTC as an alternative to complete the task.
LTU is ideal for integrating well-established domain-specific tools with specialized functionalities (for example, alignment, variant call-ing, cell clustering), which often involve complex dependencies. In fast-evolving omics research, these tools often come with complex environments and set-up procedures. We did not want BioMedAgent to be stuck solving set-up issues instead of doing the actual analysis, so we discouraged it from creating its own codes from scratch for such tasks. When specialized tools are available for the core analytical steps, and BioMedAgent only needs to handle lightweight tasks like filtering, splitting or result aggregation, the whole system becomes more stable, reproducible and suitable for real-world biomedical applications.
CTC works well for lightweight but essential operations such as scientific reasoning, summarizing, data filtering, splitting and aggregation, by using some publicly available packages already learned by LLMs. In cases where a required tool is missing, the agent attempts to generate its own CTC as an alternative to fill the gap.
This dual mechanism makes BioMedAgent both powerful and flex-ible, enabling it to support professional biomedical workflows while remaining adaptable and extensible across diverse research scenarios.
Inter-agent communication and self-evolving
To strengthen collaboration between agents, we introduced inter-agent communication and memory retention mechanisms through our IE and MR algorithms. IE allows agents, such as the planning agent and coding agent, or the coding agent and execution agent, to exchange feedback and context across steps, enabling dynamic adaptation and higher task success rates. MR assigns dedicated memory spaces to each agent (plan-ning, coding and tool agents) and uses retrieval-augmented techniques to help agents reuse past experiences, making the system self-evolving.
Automatic code repair by the IE algorithm. BioMedAgent divides the workflow into multiple steps, transforming each step into a subtask. The Programmer agent then writes code for these subtasks using existing tool libraries and generates unit test code to verify correct-ness. The Executor agent runs this test code, handling variables, files and resources while producing the corresponding outputs. A shared resource pool maintains user inputs and intermediate outputs, ensuring that each step can access the necessary data. This structured approach ensures reliable and efficient code generation in multistep analytical tasks.
Bugs and errors are inevitable in the initial programming, and even experienced human programmers must iteratively refine and improve their code to achieve correct execution. To address this chal-lenge, BioMedAgent uses a robust IE strategy for automated code repair. By combining forward information propagation and backward feedback, the IE algorithm enables BioMedAgent to broadcast localized issues to multiple upstream and downstream agents, transforming the problem-solving process from individual correction to collaborative intelligence.
When a unit test fails, the Executor agent collects detailed error information and forwardsit to the Programmer agent group for explo-ration and discussion. This group examines the errors across dimen- sions such as syntax, parameters, logic and formatting, identifying the main cause and proposing corrective actions (Supplementary Fig. 2c). Empirically, IE effectively facilitates collaborative problem-solving, enabling multiple agentstosurpass individual limitations by interact-ing and exploring together. It has successfully resolved most logical errors and code vulnerabilities introduced in the planning and coding phase. However, some issues may exceed the capabilities of LLMs, and repeated IE attempts might still lead to errors. To address this, we set an upper limit of three interaction rounds for IE during the planning and coding phases.
Tool, planning and coding memory in the MR algorithm. Memory helps organisms adapt to their environment, improve their behav-iour, and learn socially as they evolve. Similarly, in intelligent agents, memory is essential for improving individual skills and ensuring effec-tive communications71. In BioMedAgent, we include experiences from each problem we analyse and solve into three types of memories: tool memory, planning memory and coding memory.
Tool memory. When BioMedAgent first encountersa problem, it checks which tools are relevant in its workspace and figures out how to use them to solve the problem. Initially, it concentrates on each toolindi-vidually rather than how the tools work together. Once finishing the analysis and delivering the final result, BioMedAgent records each tool’s contribution to the workflow and keeps this information in its memory.
Planning memory. While tool memory improve show each tool works, planning memory increases BioMedAgent’s overall effectiveness. After completing an analysis, BioMedAgent simplifies the workflow by replacing specific filenames and unique identifiers with general placeholders, storing this simplified version in memory. This makes the workflow useful for more types of similar problems.
Coding memory. BioMedAgent also draws on previous coding experi-ences for similar programming tasks. This approach enables code reuse, reducing the cost of creating new code. After completing an analysis, BioMedAgent sets up key-value pairs that connect the goal to be achieved in each step to its specific code implementation. These pairs are saved in memory, allowing tasks to be matched based on their descriptions rather than exact code comparison, which is often more complex.
CMA and IMF strategy for memory update. In the self-evolving learn-ing process, each learning phase is considered a‘round’. Within around, experiences are accumulated independently, while memory updates occur between rounds. For BioMedAgent, we established two strategies for updating memory: CMA and IMF.
CMA. To maximize the preservation of group intelligence, this strategy retains all memories generated within a round without filtering or forgetting. This allows BioMedAgent to maintain a comprehensive memory base, storing all past experiences, which results in a large memory size.
IMF. Inspired by human memory’s natural forgetting process, this strat-egy compares the solutions used for similar problems across different rounds. It retains the path with fewer steps and discards the others, treating the simpler steps as more efficient. This approach ensures that only one memory is kept for each type of problem, providing a richer variety of memories within the same memory space.
Reproducible, scalable and shareable intelligence with BioMedAgent
BioMedAgent addresses key challenges in biomedical data analysis by optimizing tool management, workflow planning and execution. It facilitates natural-language-based task initiation, enabling direct participation from biomedical professionals without specialized computational or bioinformatics training. The IE and MR frame-work allows BioMedAgent to accumulate extensive experience from tasks submitted by various users, all stored in natural language. This approach, centred on natural language for initiating tasks and storing experiences, enhances accessibility and scalability. By ena-bling users to create, refine, and share tools and insights through an intuitive, language-driven interface, BioMedAgent would help build a biomedical data analysis community around specific topics (Supplementary Fig. 3).
BioMedAgent could be deployed locally using online backends such as GPT-4/4o/4omini, Claude3 and local open-source LLMs such as Qwen3-235b-a22b. It offers an intuitive chat interface that simplifies initiating tasks, monitoring progress, and accessing results, making complex workflows more manageable and efficient on a local scale (Supplementary Fig. 3). This local deployment strategy improves data security and supports the development of specialized biomedical analysis systems. By customizing computational tools and accumulat-ing experiences, BioMedAgent effectively adapts to the unique needs of specific tools and research domains, driving continuous advancements and fostering deeper domain-specific insights.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The BioMed-AQA benchmark (n = 327 open questions) introduced in this study is publicly available on Hugging Face athttps://hugging- face.co/datasets/BOBQWERA/biomed-aqa-dataset, which includes the full set of natural-language questions, reference steps and mile-stones for evaluation. The complementary BioMed-AQA-MCQ subset (n=172MCQs) is available at https://huggingface.co/datasets/BOBQW-ERA/biomed-mcq-dataset. The corresponding benchmark-related input data and milestone reference files are available via Zenodo at https://doi.org/10.5281/zenodo.17430550 (ref. 72). Evaluation test- ing of BioMedAgent for rounds = 0, 1, 2 under CMA and IMF on the BioMed-AQA benchmark, along with the interactive chat details that show the full process of task planning, execution and summarization for each question, are available at http://biomed.drai.cn.
Code availability
The open-source implementation of BioMedAgent, including its multi-agent framework and autoscoring evaluation agent, is available on GitHub athttps://github.com/BOBQWERA/BioMedAgent.
References
1. Agrawal, R. & Prabakaran, S. Big data in digital healthcare: lessons learnt and recommendations for general practice. Heredity 124, 525–534 (2020).
2. Shilo, S., Rossman, H. & Segal, E. Axes of a revolution challenges and promises of big data in healthcare. Nat. Med. 26, 29–38 (2020).
3. Woldemariam, M. T. & Jimma, W. Adoption of electronic health record systems to enhance the quality of healthcare in low-income countries: a systematic review. BMJ Health Care Inform. 30, e100704 (2023).
4. Liang, H. et al. Evaluation and accurate diagnoses of pediatric diseases using artificial intelligence. Nat. Med. 25, 433–438 (2019).
5. Xu, H. et al. A whole-slide foundation model for digital pathology from real-world data. Nature 630, 181–188 (2024).
6. Feinberg, D. A. et al. Next-generation MRI scanner designed for ultra-high-resolution human brain imaging at 7 Tesla. Nat. Methods 20, 2048–2057 (2023).
7. Schuijf, J. D. et al. CT imaging with ultra-high-resolution:opportunities for cardiovascular imaging in clinical practice.J. Cardiovasc. Comput. Tomogr. 16, 388–396 (2022).
8. Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
9. Metzker, M. L. Sequencing technologies—the next generation.Nat. Rev. Genet. 11, 31–46 (2010).
10. Karczewski, K. J. & Snyder, M. P. Integrative omics for health and disease. Nat. Rev. Genet. 19, 299–310 (2018).
11. Van de Sande, B. et al. Applications of single-cell RNA sequencing in drug discovery and development. Nat. Rev. Drug Discov. 22,496–520 (2023).
12. Wratten, L., Wilm, A. & Göke, J. Reproducible, scalable, and shareable analysis pipelines with bioinformatics workflow managers. Nat. Methods 18, 1161–1168 (2021).
13. Cao, Y. et al. Ensemble deep learning in bioinformatics. Nat. Mach. Intell. 2, 500–508 (2020).
14. Dubay, C. et al. Delivering bioinformatics training: bridging the gaps between computer science and biomedicine. Proc. AMIASymp.2002, 220–224 (2002).
15. Elmarakeby, H. A. et al. Biologically informed deep neural network for prostate cancer discovery. Nature 598, 348–352 (2021).
16. Li, J. et al. Proteomics and bioinformatics approaches for identification of serum biomarkers to detect breast cancer. Clin. Chem. 48, 1296–1304 (2002).
17. Sadybekov, A. V. & Katritch, V. Computational approaches streamlining drug discovery. Nature 616, 673–685 (2023).
18. Fitzgerald, R. C. et al. The future of early cancer detection. Nat. Med. 28, 666–677 (2022).
19. McDonald, T. O. et al. Computational approaches to modelling and optimizing cancer treatment. Nat. Rev. Bioeng. 1, 695–711 (2023).
20. Misra, B. B. et al. Integrated omics: tools, advances, and future approaches. J. Mol. Endocrinol. 62, R21–R45 (2019).
21. Brooks, T. G. et al. Challenges and best practices in omics benchmarking. Nat. Rev. Genet. 25, 326–339 (2024).
22. Di Tommaso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
23. Goecks, J. et al. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, 1–13 (2010).
24. Malhotra, R. et al. Using the seven bridges Cancer Genomics Cloud to access and analyze petabytes of cancer data. Curr. Protoc. Bioinformatics 60, 11.16.1–11.16.32 (2017).
25. Kaur, S. & Kaur, S. Genomics with cloud computing. Int. J. Sci. Technol. Res. 4, 146–148 (2015).
26. Wu, T. et al. A brief overview of ChatGPT: the history, status quo and potential future development. IEEE/CAAJ. Autom. Sin. 10, 1122–1136 (2023).
27. Brown, T. B. et al. Language models are few-shot learners. Adv. Neural Inf. Process. Syst. 33, 1877–1901 (2020).
28. Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
29. Hao, M. et al. Large-scale foundation model on single-cell transcriptomics. Nat. Methods 21, 1481–1491 (2024).
30. Singhal, K. et al. Large language models encode clinical knowledge. Nature 620, 172–180 (2023).
31. Tang, X. et al. MedAgents: large language models as collaborators for zero-shot medical reasoning. Find. Assoc. Comput. Linguist: ACL 2024, 599–621 (2024).
32. Guo, T. et al. Large language model based multi-agents: a survey of progress and challenges. Proc. Int. Joint Conf. Artif. Intell. 33, 8048–8057 (2024).
33. Wang, H. et al. Scientific discovery in the age of artificial intelligence. Nature 620, 47–60 (2023).
34. Gao, S. et al. Empowering biomedical discovery with AI agents. Cell 187, 6125–6151 (2024).
35. Boiko, D. A. et al. Autonomous chemical research with large language models. Nature 624, 570–578 (2023).
36. Dai, T. et al. Autonomous mobile robots for exploratory synthetic chemistry. Nature 635, 890–897 (2024).
37. Hou, W. & Ji, Z. Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis. Nat. Methods 21, 1462–1465 (2024).
38. Lobentanzer, S. et al. A platform for the biomedical application of large language models. Nat. Biotechnol. 43, 166–169 (2025).
39. Tayebi Arasteh, S. et al. Large language models streamline automated machine learning for clinical studies. Nat. Commun. 15, 1603 (2024).
40. Zhou, J. et al. An AI agent for fully automated multi-omic analyses. Adv. Sci. 11, e2407094 (2024).
41. Xiao, Y. et al. CellAgent: an LLM-driven multi-agent framework for automated single-cell data analysis. Preprint at bioRxivhttps:// doi.org/10.1101/2024.05.13.593861(2024).
42. Liu, H. & Wang, H. GenoTEX: a benchmark for evaluating LLM-based exploration of gene expression data in alignment with bioinformaticians. Preprint at arXiv https://doi.org/10.48550/ arXiv.2406.15341 (2024).
43. Mitchener, L. et al. Bixbench: a comprehensive benchmark for LLM-based agents in computational biology. Preprint at arXivhttps://doi.org/10.48550/arXiv.2503.00096(2025).
44. Gómez-López, G. et al. Precision medicine needs pioneering clinical bioinformaticians. Brief. Bioinform. 20, 752–766 (2019).
45. Hou, X. et al. Large language models for software engineering: A systematic literature review. ACM Trans. Softw. Eng. Methodol. (2023).
46. Xin, Q. et al. BioInformatics Agent (BIA): unleashing the power of large language models to reshape bioinformatics workflow. Preprint at bioRxivhttps://doi.org/10.1101/2024.05.22.595240 (2024).
47. Su, H., Long, W. & Zhang, Y. BioMaster: multi-agent system for automated bioinformatics analysis workflow. Preprint at bioRxiv https://doi.org/10.1101/2025.01.23.634608 (2025).
48. Afzal, M. et al. Precision medicine informatics: principles, prospects, and challenges. IEEE Access 8, 13593–13612 (2020).
49. Zhang, W. & Mei, H. A constructive model for collective intelligence. Natl Sci. Rev. 7, 1273–1277 (2020).
50. Qian, C. et al. Iterative experience refinement of software-developing agents. Preprint at arXiv https://doi.org/10.48550/ arXiv.2405.04219 (2024).
51. Riffle, D. et al. OLAF: an open life science analysis framework for conversational bioinformatics powered by large language models. Preprint at arXivhttps://doi.org/10.48550/ arXiv.2504.03976 (2025).
52. Xie, E. et al. CASSIA: a multi-agent large language model for automated and interpretable cell annotation. Nat. Commun. 17, 389 (2025).
53. Tsyganov, M. M. et al. Influence of DNA copy number aberrations in ABC transporter family genes on the survival of patients with primary operatable non-small cell lung cancer. Curr. Cancer Drug Targets (2025).
54. Sun, Y. et al. SERINC2-mediated serine metabolism promotes cervical cancer progression and drives T cell exhaustion. Int. J. Biol. Sci. 21, 1361–1377 (2025).
55. Wang, X., Jiang, C. & Li, Q. Serinc2 drives the progression of cervical cancer through regulating Myc pathway. Cancer Med. 13, e70296 (2024).
56. Lee, J. S. et al. SEZ6L2 is an important regulator of drug-resistant cells and tumor spheroid cells in lung adenocarcinoma. Biomedicines 8 (2020).
57. Ishikawa, N. et al. Characterization of SEZ6L2 cell-surface protein as a novel prognostic marker for lung cancer. Cancer Sci. 97, 737–745 (2006).
58. Jee, J. et al. DNA liquid biopsy-based prediction of cancer-associated venous thromboembolism. Nat. Med. 30, 2499–2507 (2024).
59. Xu, Z. et al. MiHATP: a multi-hybrid attention super-resolution network for pathological image based on transformation pool contrastive learning. In International Conference on Medical Image Computing and Computer-Assisted Intervention (Springer, 2024).
60. Yang, K. et al. If LLM is the wizard, then code is the wand: a survey on how code empowers large language models to serve as intelligent agents. Preprint at arXiv https://doi.org/10.48550/ arXiv.2401.00812(2024).
61. Qian, C. et al. Investigate-consolidate-exploit: A general strategy for inter-task agent self-evolution. Preprint at arXiv https://doi.org/10.48550/arXiv.2401.13996 (2024).
62. Gao, C. et al. Large language models empowered agent-based modeling and simulation: a survey and perspectives. Humanit. Soc. Sci. Commun. 11, 1–24 (2024).
63. Zhong, W. et al. Memorybank: enhancing large language models with long-term memory. Proc. AAAI Conf. Artif. Intell. 38, 19724– 19731 (2024).
64. Liu, L. et al. Think-in-memory: Recalling and post-thinking enable llms with long-term memory. Preprint at arXiv https://doi.org/ 10.48550/arXiv.2311.08719(2023).
65. Li, Z. et al. VarBen: generating in silico reference data sets for clinical next-generation sequencing bioinformatics pipeline evaluation. J. Mol. Diagn. 23, 285–299 (2021).
66. Pendleton, M. et al. Assembly and diploid architecture of an individual human genome via single-molecule technologies. Nat. Methods 12, 780–786 (2015).
67. Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
68. Yang, Y. et al. Comprehensive landscape of resistance mechanisms for neoadjuvant therapy in esophageal squamous cell carcinoma by single-cell transcriptomics. Signal Transduct. Target. Ther. 8, 298 (2023).
69. Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 1–5 (2018).
70. Bu, D. et al. KOBAS-i: intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res. 49, W317–W325 (2021).
71. Jiang, X. et al. Long term memory: the foundation of Al self-evolution. Preprint at arXiv https://doi.org/10.48550/ arXiv.2410.15665(2024).
72. Sun, J. Biomedical dataset files collection. Zenodo https://doi.org/ 10.5281/zenodo.17430550 (2025).
Acknowledgements
This work was supported by the National Key R&D Program of China (2022YFF1203303, D.B.), National Natural Science Foundation of China (32341019, Y.Z.; 92474204, Y.Z.; 32570778, D.B.; W2431057, K.Z.), Ningbo Top Medical and Health Research Program (2023030615, Y.Z.; 2024020919, Y.W.), Beijing Natural Science Foundation (L222007, D.B.), Ningbo Science and Technology lnnovation Yongjiang 2035 Project (2023Z226 and 2024Z229, Y.Z.), Major Project of Guangzhou National Laboratory (GZNL2023A03001, Y.Z.), State Key Laboratory of Systems Medicine for Cancer (KF2422-93, Y.Z.), Hubei Province key research and development project (2022BCA016 and 2023BCB146, Y.J.) and Macau Science and Technology Development Fund (0007/2020/AFJ, 0070/2020/A2 and 0003/2021/AKP, K.Z.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. The authors would like to acknowledge the Nanjing lnstitute of lnforSuperBahn OneAiNexus for providing the training and evaluation platform.
Author contributions
D.B. collected and interpreted the data and drafted the manuscript and figures. J.S. implemented and evaluated the algorithm and developed the software. K.L. analysed the data, drafted the figures and contributed to algorithm application. Z.H., W.H., J.H., S.Z. and S.L. participated in data processing and algorithm evaluation. P.H., Z.W. and S.W. contributed to tool preparation and website construction. T.W., K.G. and Y.W. assisted with literature collection and interpreted the results. L.Z., K.W., G.L., H.S. and Y.J. interpreted the results and supported application design. K.Z. and R.C. interpreted the results, revised the manuscript and figures and supervised the project. Y.Z. conceived the study, collected the data, revised the manuscript and figures and supervised the project. All authors reviewed and approved the final version of the manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Extended data is available for this paper at https://doi.org/10.1038/s41551-026-01634-6.
Supplementary information The online version contains supplementary material available at https://doi.org/10.1038/s41551-026-01634-6.
Correspondence and requests for materials should be addressed to Kang Zhang, Runsheng Chen or Yi Zhao.
Peer review information Nature Biomedical Engineering thanks Feixiong Cheng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Reprintsand permissions information is available at www.nature.com/reprints.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
© The Author(s), under exclusive licence to Springer Nature Limited 2026
1Research Center for Ubiquitous Computing Systems, lnstitute of Computing Technology, Chinese Academy of Sciences, Beijing, China. 2University of Chinese Academy of Sciences, Beijing, China. 3Al Cross Disciplinary Research lnstitute and Faculty of Medicine, Macau University of Science and Technology, Macau, China. 4Guangzhou National Laboratory, Guangzhou, China. 5Ningbo No.2 Hospital, Ningbo, China. 6Henan lnstitute of Advanced Technology, Zhengzhou University, Zhengzhou, China. 7Luoyang lnstitute of lnformation Technology lndustries, Luoyang, China. 8Beijing University of Chinese Medicine, Beijing, China. 9Department of Big Data and Biomedical Al, College of Future Technology, Peking University and Peking-Tsinghua Center for Life Sciences, Beijing, China. 10State Key Laboratory of Eye Health and National Clinical Research Center for Ocular Diseases, Eye Hospital, Wenzhou Medical University, Wenzhou, China. 11West China Biomedical Big Data Center, West China Hospital, Sichuan University, Chengdu, China. 12Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, China. 13Key Laboratory of RNA Biology, Center for Big Data Research in Health, lnstitute of Biophysics, Chinese Academy of Sciences, Beijing, China. 14These authors contributed equally: Dechao Bu, Jingbo Sun, Kun Li. 15These authors jointly supervised this work: Kang Zhang, Runsheng Chen, Yi Zhao. e-mail: kang.zhang@gmail.com; crs@ibp.ac.cn; biozy@ict.ac.cn










downloadFile